1) 36 pts. Short answer section.
(a) 4 pt. What is meant by a helper transposon or a helper virus?
A helper element
provides a function needed by a defective element in order to transpose or to
grow. For example enhancer trap
P-elements no longer carry transposase and require a helper to provide the
enzyme. Similarly, defective
retroviruses need a helper virus to provide the missing viral function
(b) 4 pt. Which proteins constitute the HIV receptor?
The CD4 and CCR-5 proteins together constitute the primary HIV receptor. Late in infection M-trophic viruses recognize a receptor composed of CD4 and CXCR-4.
(c) 8 pt. State whether the following best apply to E. coli lac, or yeast GAL regulation, or whether they are equally applicable.
(i) Inducer inactivates a repressor, thus allowing an activator to function.
Applies
to both. In lac, inducer inactivates LacI and CAP-cAMP can
stimulate transcription. In the
yeast GAL system, inducer
inactivates GAL80 uncovering the activation domain of GAL4 and transcription of
the GAL genes occurs.
(ii) Mutations in the activator can bypass the effect of uninducible repressor mutants.
Applies to yeast GAL.
GAL4 activators that cannot bind an uninducible version of GAL80 will
express the GAL genes. An E. coli CAP* mutant will still be repressed
by an IS
repressor.
(d) 6 pt. Predict the expression pattern
of the lac operon in an E. coli merodiploid with genotype lacIQ/lacI-d. Explain your answer.
The strain will be
inducible. lacI-d
is a dominant negative mutation
that functions as a poison subunit in the repressor tetramer leading to
constitutive expression even when wild-type subunits are present. However, lacIQ produces 10-20-times the normal amount of
wild-type repressor, meaning that the majority of repressor molecules in the IQ/I-d
merodiploid cell will be composed of wild-type subunits and therefore be
capable of binding DNA and being induced by IPTG.
(e) 8 pt. An enhancer trap screen have identified 5 independent fly lines that express ß-galactosidase in the developing trachea, or breathing tubes, of the fly. All 5 enhancer trap lines are viable when homozygous, and none cause tracheal defects. Explain what you could do to determine whether any of the enhancer trap lines affect a gene that is important for tracheal function or development.
It's possible that none
of the P insertions has inactivated the gene. To see if the genes are important for tracheal function one
needs to make loss-of-function mutations.
This can be done by screening for imprecise excisions, or
P-element-induced deletion derivatives of these genes. This can be done by exposing the flies
to a transposase source and then looking for white-eyed ß-gal non-expressors. These canb e examined by Southern blotting to see if the
gene has been deleted by imprecise excision. Once a deletion has been isolated for a particular gene,
flies can be made homozygous for the deletion and their phenotypes
examined. If the gene is
important for viability flies homozygous for the P-induced deletions will not
survive or will not develop normal trachea.
(f) 6 pt. Suggest two ways that a proto-oncogene can be converted into an active oncogene.
A protooncogene could be
activated by a point mutation such as the one that makes the Ras kinase
constitutively active.
Another way would be to
alter the expression of an oncogene.
Unregulated transcription of myc due to fusion of the gene and its
promoter to a new enhancer can cause constitutive high-level expression of this
cell-cycle activator.
2) 15 pts. The pheR100 deletion results in the constitutive expression of the phe operon (pheRABCD) of a hypothetical Bacillus. DNA from wild-type strain pheR+ and DNA from the pheR100 mutant were isolated an used in an in vitro transcription system containing only purified RNA polymerase, the DNAs, and rNTPs. The following results were obtained:
|
DNA |
relative amount of pheABCD RNA made |
|
pheR+ |
0.2 |
|
pheR100 |
10 |
a) The results of this experiment and the in vivo phenotype of pheR100 argue against the existence of a phe operon repressor. Why?
Since
there is no repressor protein in the in vitro system, there should be no
repression of either template.
Both the wild-type and the deletion pheR100 DNAs should express equally well since the
operator would be unbound in the case of wild-type or absent in the case o the
deletion.
b) The results also argue against the existence of a positive activator protein. Why?
Because
the in vitro system does not contain the activator protein, one would expect
low-level expression for both templates.
For the wild-type, the binding site is there, but unoccupied. For the mutant the site is missing and
unoccupied.
c) Based on what you know about gene regulation in E. coli, suggest what sort of site is removed by the pheR100 mutation.
It seems that something
serves as a barrier to transcription in the wild-type and that the pheR100
deletion removes the barrier. This
suggests that the site is likely a transcription terminator or an attenuator.
3) 16 pts.
The SOS DNA repair genes in E. coli are negatively regulated by the product of the lexA gene, called the LexA repressor. When a cell sustains extensive damage
to its DNA, LexA repressor is inactivated and transcription of the SOS genes is
increased dramatically.
![]()
One of the SOS genes is uvrA. You isolate a strain with constitutive
expression of uvrA protein and
name the mutation uvrA(con). Shown below is a diagram of the uvrA and lexA
genes.
(a) Describe two different mutations that would result in a uvrA constitutive phenotype.
Mutations that inactivated lexA should lead to a constitutive uvrA phenotype as the uvrA gene will no longer be repressed. Mutations in the uvrA operator should also cause a constitutive uvrA phenotype as they would block the binding of LexA.
(b) Outline an experiment that would allow you to determine which of the two mutations you have isolated.
Make a partial diploid
with the constitutive mutation and a wild-type lexA and uvrA genes. If lexA is mutated, it should be a recessive mutation
and uvrA will be repressed
and inducible by DNA damage. If
the mutation is in the uvrA
operator, uvrA will still be
constitively expressed, as LexA will not bind the mutant copy of the gene.
One could also examine
whether the other SOS genes are induced in the uvrA(con) strain. If they are it would imply that lexA is mutant, as the cis-acting uvrA operator mutation should only affect genes in
the uvrA operon. If it appears to affect lexA, one should confirm this result by showing that
the uvrA(con) allele is recessive to lexA+, to exclude the possibility that the constitutivity
arises from another mutation, i.e. one that constitively expresses the inducing
signal. Such mutations exist and
are known as recA*.
4) 33 pts.
A novel steroid hormone receptor (TaxR) has recently been identified. The receptor responds to the level of
Test Anxiety Hormone circulating in the blood and mediates a variety of
biological responses ranging from muscle twitching to complete repression of
cerebral function.
By analogy, TaxR
is suspected to contain four different protein domains: a hormone binding site, a region that
interacts with the hsp90 protein, a DNA binding domain, and a transcriptional
regulatory region. To map the TaxR
protein domains a variety of TaxR
deletion mutants were created using recombinant DNA techniques. These TaxR mutants were analyzed in cultured mammalian cells
that carried a lacZ+ reporter
gene whose expression was controlled by four TRE sequences (TRE=Test
Anxiety Hormone Receptor Element).
To monitor intracellular location, the mutant TaxR genes were fused to a gene encoding a protein called GFP (Green Fluorescent Protein). GFP is naturally fluorescent--that is, the protein glows bright green when stimulated with light.
The Tax-R derivatives
were transformed into cells and assayed with or without Test Anxiety
hormone. The results for the
wild-type protein and for a full-length TaxR-GFP fusion are shown:

The
domains of TaxR are labeled 1-4.
GFP protein is shown in gray.
Fluorescent cells and nuclei are gray. White indicates no fluorescence.
(a) 6 pts. Are the results consistent with those obtained for
other steroid hormone receptors? Briefly explain why or why not.
Yes, the TaxR protein is cytoplasmic and
inactive unless hormone is present.
In response to hormone TaxR enters the nucleus and activates
transcription. This is exactly the
effect seen with the Glucocorticoid receptor.
4b) 15 pts. For each of the TaxR-GFP proteins diagrammed below assume: i) that region 1 is the transcriptional regulatory domain, ii) that region 2 contains the DNA binding domain, iii) that region 3 interacts with hsp90 protein, and iv) that region 4 binds Test Anxiety Hormone.
Fill in the expected results for the cell fluorescence and ß-gal activity assays given the assumptions listed above. Briefly explain your answers in the space to the right of each drawing.
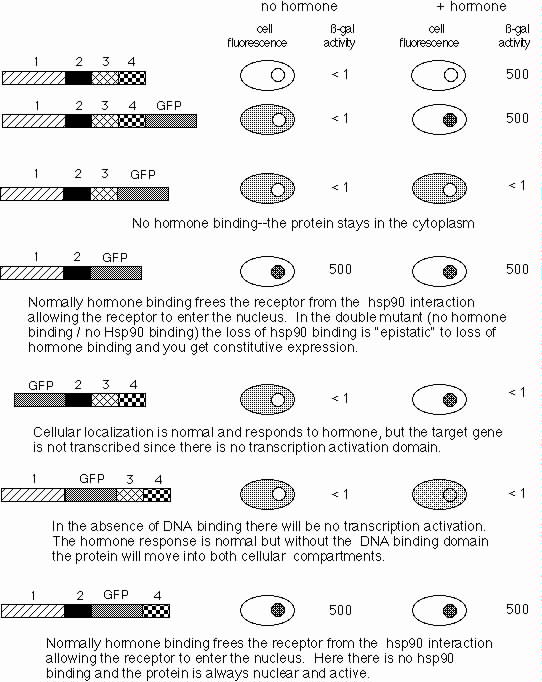
4c) 12 pts. Surprisingly TaxR appears not to interact with the hsp90 protein. Instead the data suggest it interacts with an unidentified protein thought to function analogously to hsp90.
Propose an experiment that would allow you to identify the TaxR-interacting hsp90 analog and to clone a cDNA encoding that protein. You many use any reasonable technique from the texts, the lecture, or your own experience. Your answer should provide sufficient detail to demonstrate that you understand the strategy and the techniques you employ.
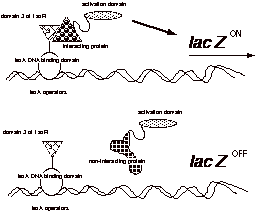
Use
the two-hybrid system to screen a cDNA library to find clones that interact
with domain 3 of TaxR. (You should
use domain 3 so that you don't recover proteins that bind to other domains of
TaxR.)
First, make a fusion between domain 3 of TaxR
and the lexA DNA binding
domain.
Second
make a random cDNA library fused to the activation domain of GAL4. The library should be made from mRNA
from tissues that are known to express the cytoplasmic interacting
protein. Given the biology of the
response, a good choice would be a library made from the brain of a student with
a minimal test anxiety response.
Introduce
the lexA-domain3 plasmid and the library plasmids into a strain carrying a lacZ gene with a minimal promoter and multiple lexA operator sites.
Select
for lacZ expression which
indicates a protein-protein interaction between domain 3 and a library
plasmid.
As
shown in the drawing, only proteins that interact with domain 3 should induce
the expression of lacZ.
To isolate the cDNA clone simply purify the strain expressing lacZ and isolate the library plasmid.
Extra credit. 6 pts. You have isolated a Tn10 insertion just upstream of the wild-type his operon following the protocol outlined in lecture. You would like to use this Tn10 to introduce a particular his- mutation (caused by a nonsense mutation in the his leader peptide) into a variety of strains with reduced tRNAHis function, ie. hisS, hisT, hisU, hisR to see what effect these mutations have on the expression of the his operon carrying the leader peptide mutation.
Explain how you would do this.
Transduce
the his- strain to
tetR. Score the tetR transductants for the his- phenotype by plating the
transductants on media with and without histidine.. Most of these will be His+ since the Tn10 is
closely linked but those that are tetR, his- now have the Tn10 tighlty linked to the his- mutation. This strain will now serve as a donor.
Grow
transducing phage on the new tetR his- donor and transduce each of the recipient strains
hisS, hisT, etc. to tetR.
Nearly all of these will carry the his- mutation.
Now
score the His phenotype for several of the resulting transductants to test for
the interaction between the altered tRNA function and the altered leader
peptide mutation.