Software Download
DTI BrainImageScope
for
Processing and Analyzing Diffusion Tensor Imaging Data
by
The DTI Laboratory, MRI Unit
Columbia University / New York State Psychiatric Institute
Supported by an NIBIB Grant 1R03EB008235-01A1 and a Shanghai CST Grant 10440710200
Contact: [email protected]
Links for Downloading:
Document:
Software:
SUN Solaris Version (still missing one module)
Test Data:
Datasets for Linux (PC-endian)
Datasets for SUN Solaris (UNIX-endian)
Brief Introduction:
This software package is developed for processing diffusion tensor imaging (DTI) data, under the auspice of National Institute of Biomedical Imaging and Bioengineering (NIBIB) and a grant from Shanghai Commission of Science and Technology.
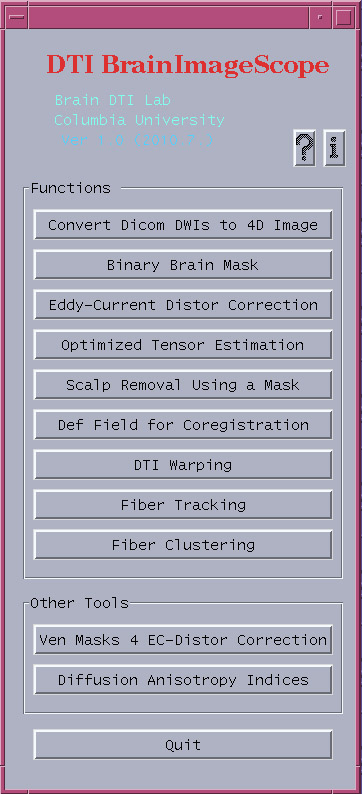
This software package is written in Python with Tkinter. The source code of the software tools and pieces integrated into this package were originally developed using C/C++, OpenGL, Matlab. Based on our work in the past years, the package covers the commonly used functions for DTI processing, including the following modules (Fig. 1):
1. Converting imaging data in DICOME format to ANALYZE format
2. Extracting binary brain mask for quick scalp-removing
3. Correcting eddy-current induced distortion
4. Optimized tensor estimation based on noisy diffusion-weighted imaging (DWI) data
5. Scalp removal using a brain mask image
6. Corregistering imaging data and generating deformation field for mapping images from individual spaces to a template or target space
7. Spatial Normalization and Warping DTI
8. Fiber tracking
9. Clustering fiber tracts
10. Identifying brain ventricles and generating binary masks for the baseline and DW imaging data
11. Deriving diffusion anisotropy indices (DAIs) and principal directions (PD) and the corresponding color-coded PD-map.

Figure 1. The main user interface of the DTI BrainImageScope Software Package.
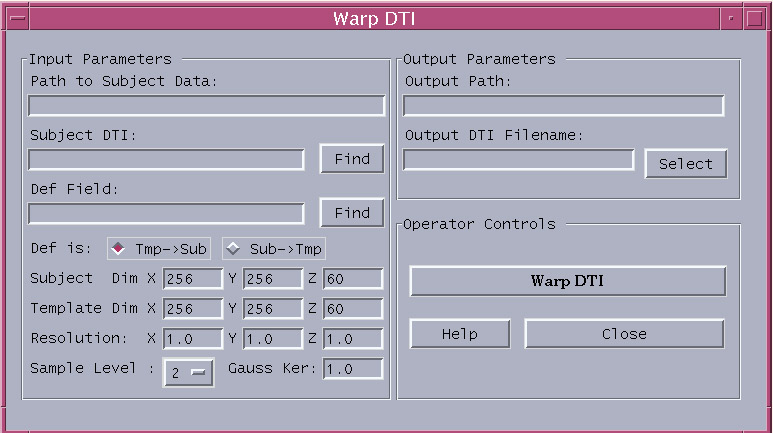
Figure 2. A typical example: The new window that will pop up if DTI Warping is clicked on the main user interface of the DTI BrainImageScope Software Package.
A Note:
The above snapshots are based on the SUN Solaris version. The
interfaces of the Linux Versions may show some misalignment, but will not affect
the functionality and execution of the functions.